New research from Kevin McKernan and Jessica Rose explains why regulators keep missing DNA contamination in mRNA vaccines—and why independent labs keep finding it.
Last September, a peer-reviewed study by genomics researcher Kevin McKernan, IMA Senior Fellow Dr. Jessica Rose, and Dr. David Speicher documented alarming levels of DNA contamination in mRNA COVID vaccines: up to 627 times above safety limits. We covered those findings here.
That study answered the question of what was in the vials. A new follow-up study, published this month in the Journal of Independent Medicine, answers an equally important question: why it was there in the first place.
Why did independent researchers keep finding contamination levels far exceeding safety limits, while regulators consistently reported acceptable levels? The answer, according to authors McKernan, Charles Rixey and Rose, comes down to enzyme chemistry—and a manufacturing decision that made contamination effectively invisible to standard testing.
📖 Read and Download the Full Paper
RNA:DNA Hybrids Survive Digestion in mRNA Vaccine Manufacturing (JIM Vol. 2, No. 1, 2026) Authors: Kevin McKernan, Charles Rixey, Jessica Rose
👉 Visit the Journal of Independent Medicine to create a free account and download the full article.
The Core Problem
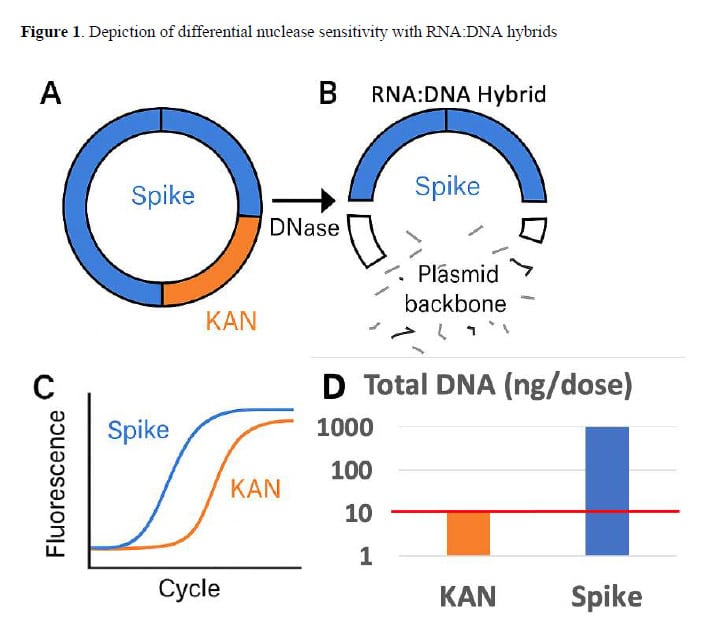
During mRNA vaccine production, DNA templates are used to generate the mRNA. That DNA is supposed to be destroyed afterward using an enzyme called DNase I. But here’s what the study demonstrates: DNase I cannot effectively digest DNA that has bonded with RNA to form what scientists call RNA:DNA hybrids.
And those hybrids form inevitably during production.
“They’re using an enzyme that’s incapable of erasing RNA-DNA hybrids,” McKernan explains. “So when they make these vaccines, they make some RNA off of the DNA, and that RNA is stuck to the DNA, and the enzymes can’t erase that RNA-DNA hybrid.”
The modifications made to the vaccine mRNA (specifically, N1-methylpseudouridine substitutions) make these bonds particularly strong and stable. Standard DNase I treatment degrades the exposed portions of the plasmid DNA but leaves these hybrid regions largely intact.
The problem: regulatory testing typically targets a region of the plasmid called the KAN (kanamycin resistance) gene—a region that is susceptible to DNase I digestion. By measuring only this region and extrapolating to estimate total DNA, regulators systematically underestimate contamination.
“This really highlights the mechanism of action and why regulators think this is below the limit and everyone else finds it above the limit,” McKernan says. “Everyone else is using tools that more comprehensively measure all of the DNA that could be there, as opposed to a really small target.”
What the Data Shows
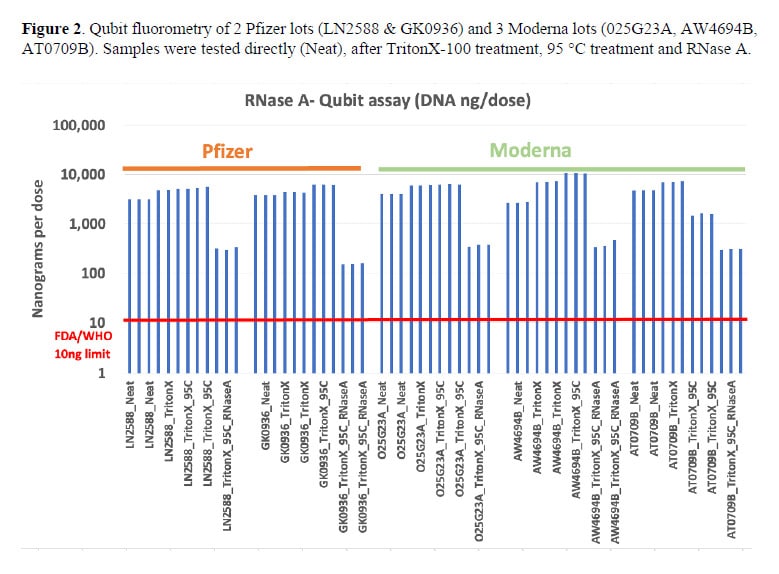
The research team analyzed two Pfizer and three Moderna vaccine lots using multiple methods: qPCR targeting different plasmid regions, fluorometry with RNase treatment, and Oxford Nanopore sequencing.
When they compared DNA measurements from the spike region (which forms hybrids) versus the origin of replication region (which doesn’t), they found measurements differing by more than 100-fold—a gap of over 7 CT values in qPCR terms.
Using fluorometry with proper controls, they detected DNA levels 15 to 48 times higher than the FDA’s recommended limit of 10 nanograms per dose. And their sequencing work recovered DNA fragments far longer than the 200 base pairs often assumed in regulatory assessments—including one fragment over 5,200 base pairs long, encompassing a large portion of the spike gene.
“When people tell you that, oh, it’s all fragmented, don’t worry about it—now we have entire coding regions in these vaccines,” McKernan notes. “Molecules that are so long that they encode the entire spike region.”
The study also tested an alternative enzyme, DNase I-XT, which is specifically designed to digest RNA:DNA hybrids. It achieved 100 to 1,000 times more effective degradation of spike-region DNA compared to standard DNase I—demonstrating that the contamination can be removed with the right tools.
Was This Known?
The mechanism the study identifies isn’t new to science. BioNTech’s own researchers published a paper in 2024 acknowledging that DNase I is at least 100-fold less effective against RNA:DNA hybrids. The appropriate enzyme exists. The problem was documented.
“There’s a citation from Lenk et al. that is actually from BioNTech themselves, publishing that they know about this,” McKernan points out. “So it’s known.”
Rose is more direct:
“I have a feeling that since we’re able to show this mechanism of action, of a lack of functionality of the right enzyme, that people did know about this. I would find it very hard to believe that they didn’t.”
What the Authors Say This Could Mean
Beyond the study’s core findings, McKernan and Rose point to a web of related concerns that they believe warrant further investigation.
They note that the commercial vaccines were manufactured using a different process than what was used during clinical trials. The trial material used PCR-generated DNA templates; the commercial product switched to E. coli-based plasmid manufacturing. This introduced not only the RNA:DNA hybrid problem but also bacterial methylation patterns.
“The vaccines that are on the market never went through clinical trial,” McKernan states. “They’re different. They’re biologically different. They’re methylated differently. They have different types of DNA present.”
They also raise questions about biological consequences. Long DNA fragments containing functional promoters could potentially express proteins if taken up by cells—which McKernan suggests may help explain reports of persistent spike protein expression in some patients.
“This helps explain potentially why we’re seeing patients out there that are still expressing spike years later,” he says. “The only way that makes sense is if these vaccines hit some sort of immunoprivileged stem cell that doesn’t get cleared out, and this DNA is sticking around.”
The study cites research linking excess DNA to myocarditis via the cGAS-STING pathway, and to oncogenesis with chronic stimulation of that same pathway. Rose references Dr. Wafik El-Deiry’s work documenting unusual patterns of aggressive cancers emerging in younger populations—a phenomenon she believes may connect to DNA contamination’s effects on tumor suppression pathways. We covered El-Deiry’s research in detail here.
Study Limitations
The authors acknowledge certain limitations. The study analyzed five vials total, and did not measure single-stranded DNA or double-stranded RNA byproducts. Oxford Nanopore sequencing has difficulty quantifying smaller fragments. These findings, while consistent with their earlier work on 32 vials, represent a focused mechanistic investigation rather than comprehensive surveillance.
The Call to Science
McKernan’s recommendation is straightforward:
“The call to action here is pull these until they clean up this manufacturing, because there are papers linking excess DNA to myocarditis… We also have papers listed in there that demonstrate this can trigger oncogenesis.”
The current 10 nanogram limit, he argues, misses the point:
“10 nanograms of a plasmid—that’s about a billion copies. It’s really not the weight of the DNA that matters. It’s the copy number.”
- For a deeper technical dive, Jessica Rose has published her own detailed breakdown of the study’s findings and implications.
- Watch Jessica Rose and Kevin McKernan discuss their study on CHD.TV
- The full study is available open-access at the Journal of Independent Medicine.
These resources are free to download—but not free to create. Our medical journal, and every system of technology and review that powers it, requires significant time and resources to build and maintain. Please support our mission to continue sharing evidence-backed medical research with the world by donating today.